Chapter 07.
The Multivariate Normal Model
본 포스팅은 First Course in Bayesian Statistical Methods를 참고하였다.
1. Multivariate Normal Desity
univariate model에 대해서 앞선 챕터에서 이야기를 많이 했지만, 사실 현실세계에서는 multivariate인 경우가 훨씬 많다.
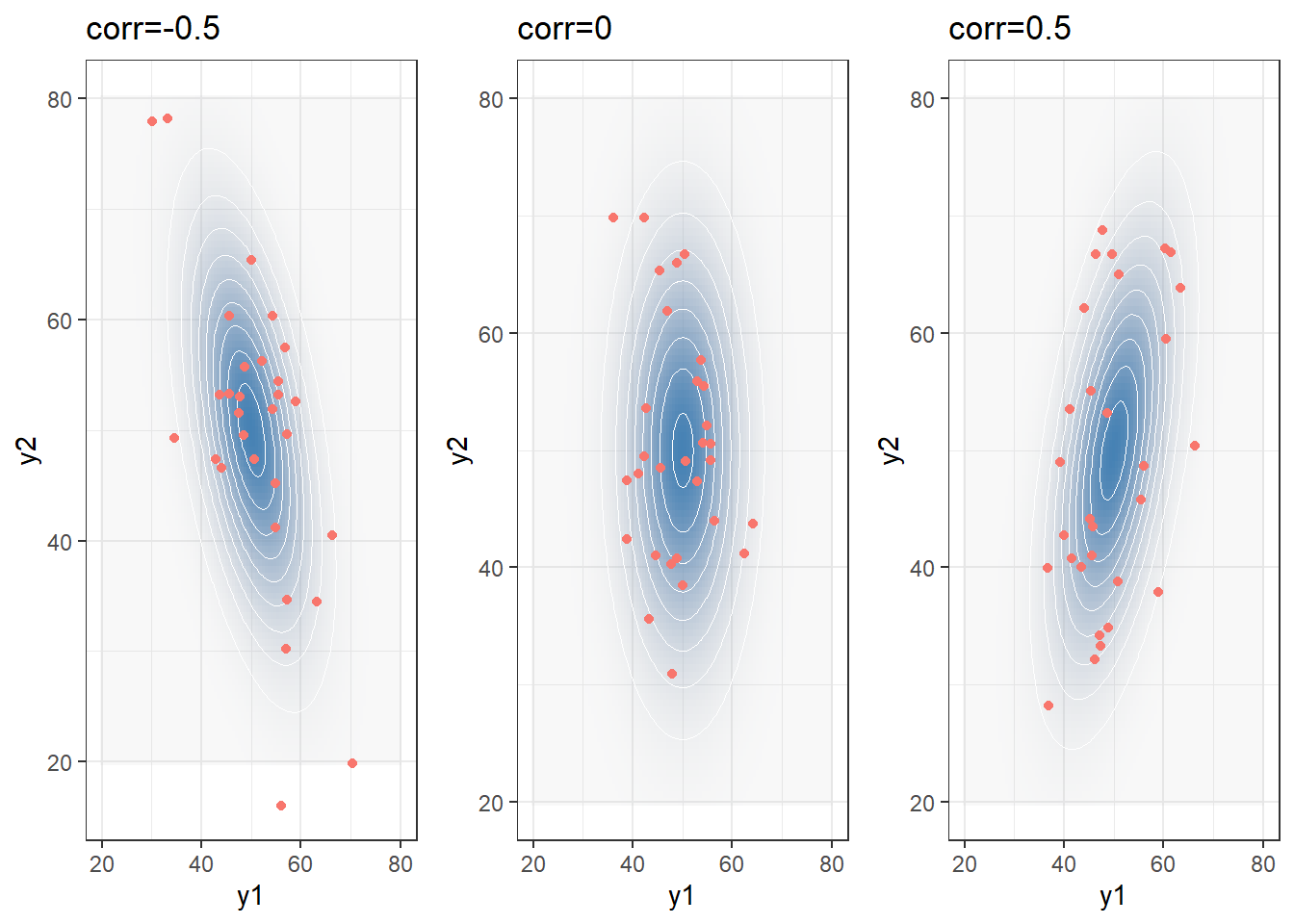
1-1. Bivariate Normal
1-2. Multivariate Normal Model
$$p(\boldsymbol{y}|\boldsymbol{\mu}, \Sigma) = (2\pi)^{-p/2}|\Sigma|^{-1/2}exp\Big(-\frac{1}{2}(\boldsymbol{y}-\boldsymbol{\mu})^T\Sigma^{-1}(\boldsymbol{y}-\boldsymbol{\mu}) \Big) $$
where
$$\boldsymbol{y} = \begin{pmatrix} y_1 \\ y_2 \\ \vdots \\ y_p \end{pmatrix}$$
$$\boldsymbol{\mu} = \begin{pmatrix} \mu_1 \\ \mu_2 \\ \vdots \\ \mu_p \end{pmatrix}$$
$$\Sigma = \begin{pmatrix} \sigma^2_{1} & \sigma_{1,2} & \cdots & \sigma_{1,p} \\ \sigma_{2,1} & \sigma^2_{2} & \cdots & \sigma_{2,p} \\ \vdots & \vdots & & \vdots \\ \sigma_{p,1} & \cdots & \cdots & \sigma^2_{p} \end{pmatrix}$$
2. Semiconjugate prior distribution for the mean (known covariance matrix)
Semiconjugate라고 하는 것은, 두 모수 중 하나가 주어졌을 경우에 conjugate한 경우를 뜻한다.
여기서는 공분산 행렬이 주어졌을 때, 평균 벡터의 semiiconjugate prior를 구하는 것(조금 더 쉬움)을 먼저 보고 이어서 공분산 행렬의 semiconjugate prior를 구하는 것을 살펴볼 것이다.
Prior: $\boldsymbol{\mu} \text{ ~ } MVN(\boldsymbol{\mu_0}, \Lambda_0)$
\begin{align} p(\boldsymbol{\mu}) &= (2\pi)^{-p/2}|\Lambda_0|^{-1/2}exp\Big(-\frac{1}{2}(\boldsymbol{\mu}-\boldsymbol{\mu_0})^T\Lambda_0^{-1}(\boldsymbol{\mu}-\boldsymbol{\mu_0})\Big) \\ &\propto exp(-\frac{1}{2}\boldsymbol{\mu}^T\Lambda^{-1}\boldsymbol{\mu} \ + \ \boldsymbol{\mu}^T\Lambda^{-1}_0\boldsymbol{\mu_0}) \\ &= exp(-\frac{1}{2}\boldsymbol{\mu}^TA_0\boldsymbol{\mu} \ + \ \boldsymbol{\mu}^T\boldsymbol{b_0}) \end{align}
where $A_0 = \Lambda^{-1}_0, \boldsymbol{b_0} = \Lambda^{-1}_0\boldsymbol{\mu_0}$
Likelihood: $Y_1, ..., Y_n|\boldsymbol{\mu},\Sigma \text{ ~ iid } MVN(\boldsymbol{\mu}, \Sigma)$
\begin{align} p(\boldsymbol{y_1}, ...,\boldsymbol{y_n}|\boldsymbol{\mu},\Sigma) &= \prod^{n}_{i=1} (2\pi)^{-p/2}|\Sigma|^{-1/2}exp\Big(-\frac{1}{2}(\boldsymbol{y_i}-\boldsymbol{\mu})^T\Sigma^{-1}(\boldsymbol{y_i}-\boldsymbol{\mu}) \Big) \\ &= (2\pi)^{-np/2}|\Sigma|^{-n/2}exp\Big(-\frac{1}{2}\sum_{i=1}^{n}(\boldsymbol{y_i}-\boldsymbol{\mu})^T\Sigma^{-1}(\boldsymbol{y_i}-\boldsymbol{\mu}) \Big) \\ &\propto exp(-\frac{1}{2}\boldsymbol{\mu}^TA_1\boldsymbol{\mu} \ + \ \boldsymbol{\mu}^T\boldsymbol{b_1}) \end{align}
where $A_1 = n\Sigma^{-1}, \boldsymbol{b_1} = n\Sigma^{-1}\boldsymbol{\bar{y}}$
Posterior: $\boldsymbol{\mu}|\boldsymbol{y_1}, ..., \boldsymbol{y_n}, \Sigma \text{ ~ } MVN(\boldsymbol{\mu_n}, \Lambda_n)$
$$
\begin{align} p(\boldsymbol{\mu}|\boldsymbol{y_1}, ..., \boldsymbol{y_n}, \Sigma) &\propto exp(-\frac{1}{2}\boldsymbol{\mu}^TA_0\boldsymbol{\mu} \ + \ \boldsymbol{\mu}^T\boldsymbol{b_0}) \times exp(-\frac{1}{2}\boldsymbol{\mu}^TA_1\boldsymbol{\mu} \ + \ \boldsymbol{\mu}^T\boldsymbol{b_1}) \\ &= exp(-\frac{1}{2}\boldsymbol{\mu}^TA_n\boldsymbol{\mu} \ + \ \boldsymbol{\mu}^T\boldsymbol{b_n}) \\ \\ \text{where } A_n &= A_0 + A_1 = \Lambda_0^{-1}+n\Sigma^{-1} \\ \boldsymbol{b_n} &= \boldsymbol{b_0} + \boldsymbol{b_1} = \Lambda_0^{-1}\boldsymbol{\mu_0}+n\Sigma^{-1}\boldsymbol{\bar{y}} \\ \\ \rightarrow \Lambda_n^{-1} &= \Lambda_0^{-1}+n\Sigma^{-1} \\ \boldsymbol{\mu_n} &= (\Lambda_0^{-1}+n\Sigma^{-1})^{-1}(\Lambda_0^{-1}\boldsymbol{\mu_0}+n\Sigma^{-1}\boldsymbol{\bar{y}}) \end{align}
$$
3. Semiconjugate prior distribution for the covariance matrix (known mean)
이는 상대적으로 복잡한데, 그 이유는 이전과 달리 matrix 형태에다가 prior을 주어야하기 때문이다.
그래서 구체적인 prior와 likelihood를 이야기하기 앞서서 필요한 두 가지 개념에 대해서 짚고 넘어가도록 하자.
첫 번째는 inverse-Wishart 분포이며, 다음은 Positive Definite이라는 선형대수 개념이다.
3-1. inverse-Wishart Distribution
inverse-Wishart Distribution은 공분산 행렬의 semiconjugate prior을 주기 위해서 알아야 하는 함수이다. 낯선 확률분포처럼 보이기도 하지만, 자세히 살펴보면 이는 inverse-Gamma distribution의 다차원 확장 버전에 불과하긴 하다.
3-2. Positive Definite
Covariance Matrix는 Positive Definite Matrix이어야 한다. Positive Definite의 정의는 아래와 같다.
$$\boldsymbol{x'}\Sigma\boldsymbol{x} > 0 \ \text{ for all vectors} \ \boldsymbol{x}$$
univariate case에서 분산이 언제나 0 이상이어야 하는 것처럼, 이와 같은 맥락의 조건을 다차원에서 만족하려면 PD(Positive Definite)이어야 한다.
만약 공분산 행렬이 Positive Definite하다면, 이는 모든 분산이 0보다 크며 공분산이 -1과 1 사이에 있도록 한다.
또한, PD는 대칭행렬(symmetric)에서 정의되는 개념이기 때문에, $\sigma_{i,j} = \sigma_{j,i}$라는 조건도 자연스럽게 성립한다.
3-2-1. Positive Definite이 되기 위한 조건은?
다시 한번 말하자면, Positive Definite은 대칭행렬의 특수한 형태이며, 모든 eigenvalue들이 0보다 크다는 말과 같다.
여기서 eigenvalue가 0보다 크다는 것은 정확히 무슨 의미일까?
정방행렬을 Spectral Decomposition을 했을 때, $A = VDV^{-1}$
$$A_{p\text{ x }p} = \begin{bmatrix} | & & |\\ a_1 & \cdots & a_p\\ | & & |\\ \end{bmatrix} = \begin{bmatrix} | & & |\\ v_1 & \cdots & v_p\\ | & & | \\ \end{bmatrix} \begin{bmatrix} \lambda_1 & & \\ & \ddots & \\ & & \lambda_p \end{bmatrix} \begin{bmatrix} | & & |\\ v_1 & \cdots & v_p\\ | & & |\\ \end{bmatrix}^{-1}$$
eigenvalue가 0보다 크다는 것은, 선형변환을 했을 때 그 기저의 방향이 반대로 바뀌지는 않는다는 것을 의미한다.
3-3. random Covariance Matrix 만들기
이는 covariance matrix에 대해 uninformative prior를 주기 위함이다.
$$\frac{1}{n}\sum_{i=1}^n\boldsymbol{z_iz_i^T} = \frac{1}{n}Z^TZ$$
$$\boldsymbol{z_iz_i^T} = \begin{pmatrix} z_{i,1}^2 & z_{i,1}z_{i,2} & \cdots & z_{i,1}z_{i,p} \\ z_{i,2}z_{i,1} & z_{i,2}^2 & \cdots & z_{i,2}z_{i,p} \\ \vdots & & & \vdots \\ z_{i,p}z_{i,1} & z_{i,p}z_{i,2} & \cdots & z_{i,p}^2 \end{pmatrix}$$
\begin{align} \frac{1}{n}\big[Z^TZ\big]_{j,j} &= \frac{1}{n}\sum_{i=1}^{n}z_{i,j}^2 = s_{j,j} = s_j^2\\ \frac{1}{n}\big[Z^TZ\big]_{j,k} &= \frac{1}{n}\sum_{i=1}^{n}z_{i,j}z_{i,k} = s_{j,k} \end{align}
여기서 n > p 이고, 모든 $\boldsymbol{z_i}$들이 서로 선형독립이라면, $Z^TZ$는 항상 positive definite일 것이다.
$$\text{Proof) } \boldsymbol{x}^{T}Z^{T}Z\boldsymbol{x} = (Z\boldsymbol{x})^{T}(Z\boldsymbol{x}) = ||Z\boldsymbol{x}||^2 \ge 0$$
STEP1. Set $\nu_0$(prior sample size), $\Phi_0$(prior covariance matrix)
STEP2. Sample $\boldsymbol{z_i} \ \stackrel{iid}{\sim} \ MVN(\boldsymbol{0}, \Phi_0)$
STEP3. Calculate $Z_TZ = \sum_{i=1}^{\nu_0}\boldsymbol{z_iz_i^T}$
STEP4. repeat the procedure S times generating $Z_i^TZ_i$
${Z_1^TZ_1, Z_2^TZ_2, ..., Z_S^TZ_S} \text{ ~ } Wis(\nu_0, \Phi_0)$
Wishart분포와 inv-Wishart분포 특징
$$\Sigma^{-1} \sim Wis(\nu_0, S_0^{-1}) \rightarrow E[\Sigma^{-1}]=\nu_0S_0^{-1} \\ \Sigma \sim Wis^{-1}(\nu_0, S_0^{-1}) \rightarrow E[\Sigma]=\frac{1}{\nu_0-p-1}S_0$$
covariance matrix semiconjugate prior 모수 설정 방법
1-1. If belief that $\Sigma = \Sigma_0$ is strong, $\nu_0$ \uparrow
1-2. If belief that $\Sigma = \Sigma_0$ is weak, $\nu_0 = p+2$
2. Set $S_0=(\nu_0-p-1)\Sigma_0$
3-4. Full conditional distribution of Covariance Matrix
Prior: $\Sigma \sim \text{inv-}Wis(\nu_0, S_0^{-1})$
$$p(\Sigma) = \bigg[2^{\nu_0p/2}\pi^{p/2}|S_0|^{\nu_0/2}\prod_{j=1}^{p}\Gamma(\frac{\nu_0+1-j}{2})\bigg]^{-1} \times |\Sigma|^{-(\nu_0+p+1)/2} \times exp\Big(-\frac{1}{2}tr(S_0\Sigma^{-1})\Big) $$
Likelihood: $\boldsymbol{Y}|\boldsymbol{\mu} \stackrel{iid}\sim MVN(\boldsymbol{\mu}, \Sigma)$
\begin{align} p(\boldsymbol{y_1, ..., y_n}|\boldsymbol{\mu}, \Sigma) &= (2\pi)^{-np/2}|\Sigma|^{-n/2} exp\bigg(-\frac{1}{2}\sum_{i=1}^{n} \boldsymbol{(y_i-\mu)}^T\Sigma^{-1}\boldsymbol{(y_i-\mu)} \bigg) \\ &\propto |\Sigma|^{-n/2}exp\bigg(-\frac{1}{2}tr(S_\mu\Sigma^{-1})\bigg) \end{align}
where $S_\mu = \sum_{i=1}^{n}\boldsymbol{(y_i-\mu)}\boldsymbol{(y_i-\mu)}^T$
Posterior: $\Sigma|\boldsymbol{y} \sim \text{inv-}Wis(\nu_0+n, [S_0+S_\mu]^{-1})$
\begin{align} p(\Sigma|\boldsymbol{y_1, ..., y_n, \mu}) &\propto p(\Sigma)p(\boldsymbol{y_1, ..., y_n}|\boldsymbol{\mu},\Sigma) \\ &\propto|\Sigma|^{-(\nu_0+p+1)/2} \times exp\Big(-\frac{1}{2}tr(S_0\Sigma^{-1})\Big) \times |\Sigma|^{-n/2}exp\bigg(-\frac{1}{2}tr(S_\mu\Sigma^{-1})\bigg) \\ &\propto |\Sigma|^{-(\nu_0+p+n+1)/2}exp\bigg(-\frac{1}{2}tr([S_0+S_\mu]\Sigma^{-1})\bigg) \end{align}
\begin{align} E[\Sigma|\boldsymbol{y_1, ..., y_n, \mu}] &= \frac{1}{\nu_0+n-p-1}(S_0+S_\mu) \\ &= \frac{\nu_0-p-1}{\nu_0+n-p-1}\cdot\frac{1}{\nu_0-p-1}S_0 + \frac{n}{\nu_0+n-p-1}\cdot\frac{1}{n}S_\mu \\ &= \frac{\nu_0-p-1}{\nu_0+n-p-1}\cdot\Sigma_0 + \frac{n}{\nu_0+n-p-1}\cdot\frac{1}{n}S_\mu \end{align}
Summary
- Semiconjugate prior for
$\mu$
Prior:$\boldsymbol{\mu} \text{ ~ } MVN(\boldsymbol{\mu_0}, \Lambda_0)$
Likelihood:$Y_1, ..., Y_n|\boldsymbol{\mu},\Sigma \text{ ~ iid } MVN(\boldsymbol{\mu}, \Sigma)$
Posterior:$\boldsymbol{\mu}|\boldsymbol{y_1}, ..., \boldsymbol{y_n}, \Sigma \text{ ~ } MVN(\boldsymbol{\mu_n}, \Lambda_n)$
\begin{align} \Lambda_n^{-1} &= \Lambda_0^{-1}+n\Sigma^{-1} \\ \boldsymbol{\mu_n} &= (\Lambda_0^{-1}+n\Sigma^{-1})^{-1}(\Lambda_0^{-1}\boldsymbol{\mu_0}+n\Sigma^{-1}\boldsymbol{\bar{y}}) \end{align}
- Semiconjugate prior for
$\Sigma$
Prior:$\Sigma \sim \text{inv-}Wis(\nu_0, S_0^{-1}) \text{ where } S_0 = (\nu_0-p-1)\Sigma_0$
Likelihood:$\boldsymbol{Y}|\boldsymbol{\mu} \stackrel{iid}\sim MVN(\boldsymbol{\mu}, \Sigma)$
Posterior:$\Sigma|\boldsymbol{y_1, ..., y_n} \sim \text{inv-}Wis(\nu_0+n, [S_0+S_\mu]^{-1})$
$$E[\Sigma|\boldsymbol{y_1, ..., y_n, \mu}] = \frac{\nu_0-p-1}{\nu_0+n-p-1}\cdot\Sigma_0 + \frac{n}{\nu_0+n-p-1}\cdot\frac{1}{n}S_\mu$$
4. Gibbs Sampling of the mean and covariance
Gibbs sampling은 full conditional distribution을 통해 차례대로 모수를 업데이트하면서 joint posterior distribution을 구하는 것이 목적이다.
STEP1. Full conditional distribution을 확보한다.
$\boldsymbol{\mu}|\boldsymbol{y_1}, ..., \boldsymbol{y_n}, \Sigma \sim MVN(\boldsymbol{\mu_n}, \Lambda_n)$$\Sigma|\boldsymbol{y_1, ..., y_n}, \boldsymbol{\mu} \sim \text{inv-}Wis(\nu_0+n, [S_0+S_\mu]^{-1})$
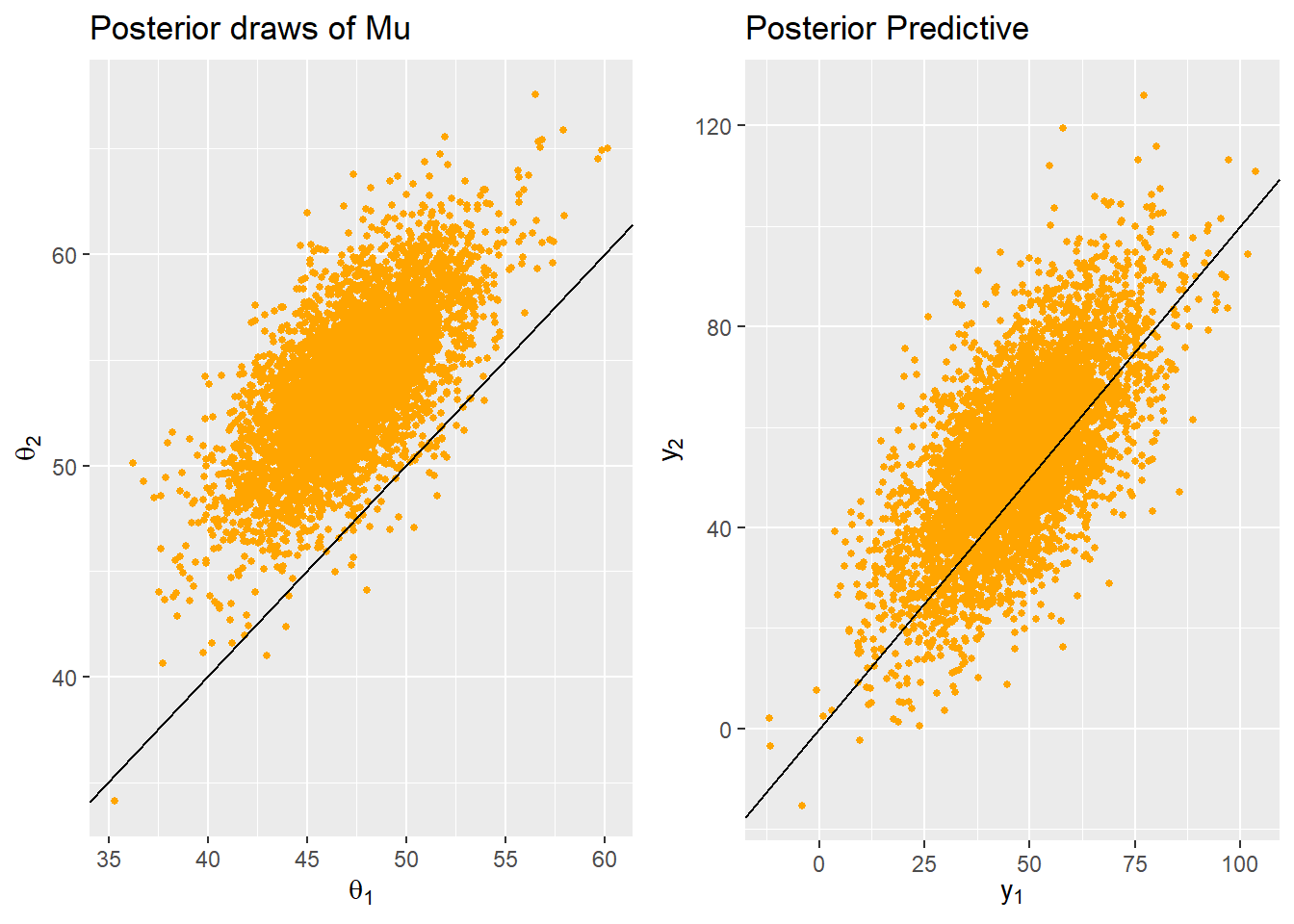
STEP2. 차례대로 업데이트하면서 joint posterior distribution $\boldsymbol{\mu},\Sigma|\boldsymbol{y_1}, ..., \boldsymbol{y_n}$을 구한다.
4-1. NA imputation
MAR(Missing at Random)인 경우에, missing data를 일종의 모수로 보고 gibbs sampling을 통해 na imputation을 해줄 수 있다.
Conclusion
MVN도 잘 알아두자. inv-Wishart 분포도!
혹시 궁금한 점이나 잘못된 내용이 있다면, 댓글로 알려주시면 적극 반영하도록 하겠습니다.